Main Page: Difference between revisions
No edit summary |
|||
| Line 13: | Line 13: | ||
= [[NgsAdmix]] = | = [[NgsAdmix]] = | ||
[[File:NgsAdmix.png|thumb]] | [[File:NgsAdmix.png|thumb|NGSadmix]] | ||
Infer the ancestry proportions from low depth NGS data. The principal is the same as other softwares such as FRAPPE and ADMIXTURE however, ngsAdmix also works when you have uncertainty in your data. This makes it ideal for medium and low depth sequencing data where many genotypes cannot be called without introducing errors or ascertainment bias. | Infer the ancestry proportions from low depth NGS data. The principal is the same as other softwares such as FRAPPE and ADMIXTURE however, ngsAdmix also works when you have uncertainty in your data. This makes it ideal for medium and low depth sequencing data where many genotypes cannot be called without introducing errors or ascertainment bias. | ||
Revision as of 11:38, 15 August 2013
ANGSD
Analysis of Next Generation Sequencing Data
<classdiagram type="dir:LR"> [sequence data]->[genotype;likelihoods] [genotype;likelihoods]->[genotype;probabilities] [sequence files|bam files;SOAP files{bg:orange}]->[sequence data] [glf files|glfv3;soapSNP{bg:orange}]->[genotype;likelihoods] [genotype prob|beagle output{bg:orange}]->[genotype;probabilities] </classdiagram>
NgsAdmix
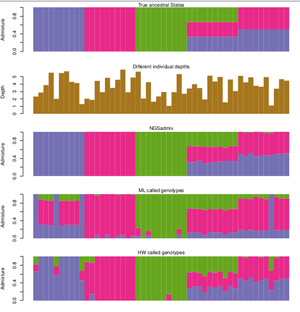
Infer the ancestry proportions from low depth NGS data. The principal is the same as other softwares such as FRAPPE and ADMIXTURE however, ngsAdmix also works when you have uncertainty in your data. This makes it ideal for medium and low depth sequencing data where many genotypes cannot be called without introducing errors or ascertainment bias.
Relate
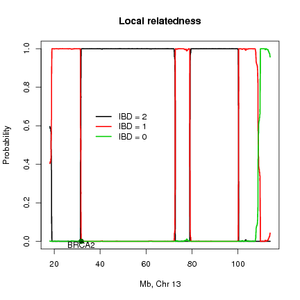
This method estimates the probability of sharing alleles identity by descent (IBD) across the genome and can also be used for mapping disease loci using distantly related individuals. To accommodate LD the methods need SNP for several individuals in order to estimate the allele frequencies and the pairwise LD. The method return the posterior probabilities of the IBD states across the genome and the overall IBD sharing.