ANGSD: Analysis of next generation Sequencing Data
Latest tar.gz version is (0.938/0.939 on github), see Change_log for changes, and download it here.
Abbababa2: Difference between revisions
No edit summary |
No edit summary |
||
| (10 intermediate revisions by 2 users not shown) | |||
| Line 1: | Line 1: | ||
The ''ABBABABA (multipop)'' compute the abbababa test (aka D-statistic), that means testing for an ancient admixture (or wrong tree topology). | The ''ABBABABA (multipop)'' compute the abbababa test (aka D-statistic), that means testing for an ancient admixture (or wrong tree topology). | ||
Differently from ABBABABA (D_stat) multiple individuals for each one of the groups are allowed. As all methods in ANGSD we require that the header of the BAM files are the same. | Differently from ABBABABA (D_stat) multiple individuals for each one of the groups are allowed. As all methods in ANGSD we require that the header of the BAM files are the same. | ||
This method [http://www.g3journal.org/content/8/2/551 has a publication]. | |||
; some of the options only works for the developmental version availeble from github | ; some of the options only works for the developmental version availeble from github | ||
; if you use -rf to specify regions. These MUST appear in the same ordering as your fai file. | ; if you use -rf to specify regions. These MUST appear in the same ordering as your fai file. | ||
| Line 34: | Line 37: | ||
-enhance 0 only analyze sites where outgroup H4 is non poly | -enhance 0 only analyze sites where outgroup H4 is non poly | ||
-Aanc 0 set H4 outgroup allele as A in each site | -Aanc 0 set H4 outgroup allele as A in each site | ||
-useLast 0 | -useLast 0 1=use the last group of bam files as outgroup | ||
</pre> | </pre> | ||
This function will counts the number of ABBA and BABA sites. | This function will counts the number of ABBA and BABA sites of the 4-population trees that can be built from the data, where the outgroup is fixed. | ||
=Output= | =Output= | ||
;1)*.abbbababa2 (used for the 4-population test) | ;1)*.abbbababa2 (used for the 4-population test) | ||
Each line represents a block with a chromsome name (Column 1), a start position (Column 2), an end postion (Column 3). The | Each line represents a block with a chromsome name (Column 1), a start position (Column 2), an end postion (Column 3). Columns 4 and 5 are the numerator and denominator of the D-statistic for their specific block. Column 6 is the number of sites containing data in that block. The other 256 columns are the normalized counts of the 256 allele patterns between the 4 populations, starting from X0000=AAAA,X0001=AAAC,....,X3333=TTTT, with the correspondence 0=A,1=C,2=G,3=T. Every block is repeated a number of times corresponding to the trees that are built. | ||
This file is used as input for the R script estAvgError.R. | This file is used as input for the R script estAvgError.R. | ||
| Line 53: | Line 56: | ||
Include an outgroup in fasta format. | Include an outgroup in fasta format. | ||
; -useLast [int] | ; -useLast [int] | ||
1: | 1: use the last group of bam files as outgroup for the D-stat analysys. Default: 0 (use the fasta file as outgroup) | ||
; -doCounts 1 | ; -doCounts 1 | ||
use -doCounts 1 in order to count the bases at each sites after filters. | use -doCounts 1 in order to count the bases at each sites after filters. | ||
| Line 63: | Line 66: | ||
allows for a maximum depth in each site to avoid overflow of the ABBA BABA counts. Default 100. | allows for a maximum depth in each site to avoid overflow of the ABBA BABA counts. Default 100. | ||
; -sizeFile [fileName] | ; -sizeFile [fileName] | ||
file that specifies number of individuals in each population (more than 4 populations can be defined) | file that specifies number of individuals in each population (more than 4 populations can be defined). If not provided, it is assumed that each population has only one individual. | ||
; -Aanc [int] | ; -Aanc [int] | ||
1: H4 allele is A in each site. | 1: H4 allele is A in each site. | ||
| Line 70: | Line 73: | ||
=Tutorial for the ABBABABA (Multipop) test= | =Tutorial for the ABBABABA (Multipop) test= | ||
This tutorial require having Samtools previously installed, and the library 'pracma' previously installed in R. | |||
== Prepare BAM and FASTA files == | == Prepare BAM and FASTA files == | ||
Download the latest version of angsd in your working folder from the github repository | Download the latest version of angsd in your working folder from the github repository | ||
| Line 104: | Line 108: | ||
bams/smallNA11840.mapped.ILLUMINA.bwa.CEU.low_coverage.20111114.bam | bams/smallNA11840.mapped.ILLUMINA.bwa.CEU.low_coverage.20111114.bam | ||
</pre> | </pre> | ||
Download a fasta file for the chimpanzee. This is going to be used as the outgroup for the 4-population test | Download a fasta file for the chimpanzee. This is going to be used as the outgroup for the 4-population test. | ||
<pre> | <pre> | ||
wget http://popgen.dk/software/download/angsd/hg19ancNoChr.fa.gz | wget http://popgen.dk/software/download/angsd/hg19ancNoChr.fa.gz | ||
| Line 110: | Line 114: | ||
gunzip chimpHg19.fa.gz | gunzip chimpHg19.fa.gz | ||
samtools faidx chimpHg19.fa #indexing the fasta file | samtools faidx chimpHg19.fa #indexing the fasta file | ||
</pre> | </pre> | ||
Now, generate a fasta file for one of our 10 bam file. We assume such a genome has very high quality and we can use it as a reference for estimating error rates in others of our datasets. | Now, generate a fasta file for one of our 10 bam file. We assume such a genome has very high quality and we can use it as a reference for estimating error rates in others of our datasets. | ||
| Line 117: | Line 120: | ||
gunzip perfectSampleCEU.fa.gz | gunzip perfectSampleCEU.fa.gz | ||
samtools faidx perfectSampleCEU.fa | samtools faidx perfectSampleCEU.fa | ||
</pre> | </pre> | ||
| Line 139: | Line 134: | ||
==4-population test== | ==4-population test== | ||
In this tutorial we perform the ABBABABA test on all the combinations of 4 populations amongst 6 populations of size 1,2,2,3,2,1 individuals, where the last population is fixed as outgroup (so that there are 30 possible combinations). The last population is represented by the fasta file defined with the option -anc, of which we enable the use as outgroup by the option -useLast 0. One can use the last population of .bam files as outgroup with the option -useLast 1. Create a file named sizeFile.size and write the size of each population (IT IS NECESSARY to define the size of the -anc outgroup population, that is always 1): | |||
<pre> | |||
1 | |||
2 | |||
2 | |||
3 | |||
2 | |||
1 | |||
</pre> | |||
We decide to target three chromosomes, one of the three with loci between position 10Mb and 15Mb. Thus create a file called regions.txt in which is written | |||
<pre> | <pre> | ||
1: | 1: | ||
5: | |||
16:10000000-15000000 | |||
</pre> | </pre> | ||
After running ANGSD we will call the R script who applies error correction to the ABBA and BABA allele combinations and produces the final output files. | The output of ANGSD will show no data about chromosome 1. This happens when all blocks within that chromosome contained no data and therefore where not printed. | ||
After running ANGSD to count ABBA and BABA combinations, we will call the R script who applies error correction to the ABBA and BABA allele combinations and produces the final output files. | |||
<pre> | <pre> | ||
./ANGSD -doAbbababa2 1 -bam bam.filelist -sizeFile sizeFile.size -doCounts 1 -out bam.Angsd -anc chimpHg19.fa -rf regions.txt -useLast | ./ANGSD -doAbbababa2 1 -bam bam.filelist -sizeFile sizeFile.size -doCounts 1 -out bam.Angsd -anc chimpHg19.fa -rf regions.txt -useLast 0 -minQ 20 -minMapQ 30 -p 1 | ||
</pre> | </pre> | ||
[[The output file is]] | [[The output file is]] | ||
[[bam.Angsd.abbbababa2 (used for the 4-population test)]] | [[bam.Angsd.abbbababa2 (used for the 4-population test)]] | ||
Each line represents a block with a chromsome name (Column 1) for one of the possible | Each line represents a block with a chromsome name (Column 1) for one of the possible 30 trees (so each block is written on 30 lines), a start position (Column 2), an end postion (Column 3). Columns 4,5 and 6 are the numerator, denominator and number of sites analyzed in the block. The next 256 columns are the counted patterns of alleles in the tree, starting from X0000=AAAA,X0001=AAAC,....,X3333=TTTT, with the correspondence 0=A,1=C,2=G,3=T. | ||
This file is used as input for the R script estAvgError.R. | This file is used as input for the R script estAvgError.R. | ||
We run the R script specifying the error files for the population with 3 individuals. This is done defining the error files in each populations inside a text file. If a population has no error file, it is sufficient to write NA. Create a file called errorList.error with written | We run the R script specifying the error files for the population with 3 individuals. This is done defining the error files in each populations inside a text file (including a line for the outgroup population). If a population has no error file, it is sufficient to write NA. Create a file called errorList.error with written | ||
<pre> | <pre> | ||
NA | NA | ||
| Line 164: | Line 169: | ||
NA | NA | ||
</pre> | </pre> | ||
Create a file popNames.name with written | |||
<pre> | <pre> | ||
Population1 | Population1 | ||
| Line 175: | Line 180: | ||
Run the Rscript with the command | Run the Rscript with the command | ||
<pre> | <pre> | ||
Rscript DSTAT angsdFile="bam.Angsd" out="result" sizeFile=sizeFile.size errFile=errorList.error nameFile=popNames.name | Rscript DSTAT angsdFile="bam.Angsd" out="result" sizeFile=sizeFile.size errFile=errorList.error nameFile=popNames.name | ||
</pre> | </pre> | ||
The script will show the calculated D statistic along with Z-score, Pvalues, Standard deviation and other quantities for all | The script will show the calculated D statistic along with Z-score, Pvalues, Standard deviation and other quantities for all 30 4-populations trees. Note: If error correction is not needed, it is sufficient to avoid specifying any error file. If no names need to be provided, the script will assign Population_* as standard name. If no size file is provided, the script assigns 1 to each population. At least one between the name file and the size file is needed. It is possible to recycle the size file used in ANGSD. | ||
The D-statistics are contained in four distinct files | The D-statistics and other informations are contained in four distinct files depending on the application of error correction and ancient transition removal. The files are named as follow: | ||
;[[1)result | ;[[1)result.Observed.txt]] | ||
D-statistic calculated WITHOUT Error Correction and WITHOUT Ancient Transition removal | D-statistic calculated WITHOUT Error Correction and WITHOUT Ancient Transition removal | ||
;[[2) result | ;[[2) result.ErrorCorr.txt]] | ||
D-statistic calculated WITH Error Correction and WITHOUT Ancient Transition removal | D-statistic calculated WITH Error Correction and WITHOUT Ancient Transition removal | ||
;[[3) result. | ;[[3) result.ErrorCorr.TransRem.txt]] | ||
D-statistic calculated WITH Error Correction and WITH Ancient Transition removal | D-statistic calculated WITH Error Correction and WITH Ancient Transition removal | ||
;[[4) result. | ;[[4) result.TransRem.txt]] | ||
D-statistic calculated WITHOUT Error Correction and WITH Ancient Transition removal | D-statistic calculated WITHOUT Error Correction and WITH Ancient Transition removal | ||
Specifically, the values contained in the four files are: mean(D)=average D-stat, JK-D=jackknife estimate of the D-stat, V(JK-D)=variance of the D-stat, Z=Z score, pvalue=pvalue from the Z score, nABBA=number of ABBA patterns observed, nBABA=number of BABA patterns observed, the names of the four populations. Note that the number of patterns might not be integer because of how ANGSD treats multiple genomes per populations. | Specifically, the values contained in the four files are: mean(D)=average D-stat, JK-D=jackknife estimate of the D-stat, V(JK-D)=variance of the D-stat, Z=Z score, pvalue=pvalue from the Z score, nABBA=number of ABBA patterns observed, nBABA=number of BABA patterns observed, nBlocks=number of blocks with observed data, H*=the names of the four populations for the specific tree. Note that the number of patterns might not be integer because of how ANGSD treats multiple genomes per populations. | ||
=Cite the method= | |||
<pre> | |||
@article{Soraggi2018, | |||
author = {Soraggi, S. and Wiuf, C. and Albrechtsen, A.}, | |||
doi = {10.1534/g3.117.300192}, | |||
issn = {21601836}, | |||
journal = {G3: Genes, Genomes, Genetics}, | |||
number = {2}, | |||
title = {{Powerful inference with the D-statistic on low-coverage whole-genome data}}, | |||
volume = {8}, | |||
year = {2018} | |||
} | |||
</pre> | |||
Latest revision as of 11:34, 26 June 2018
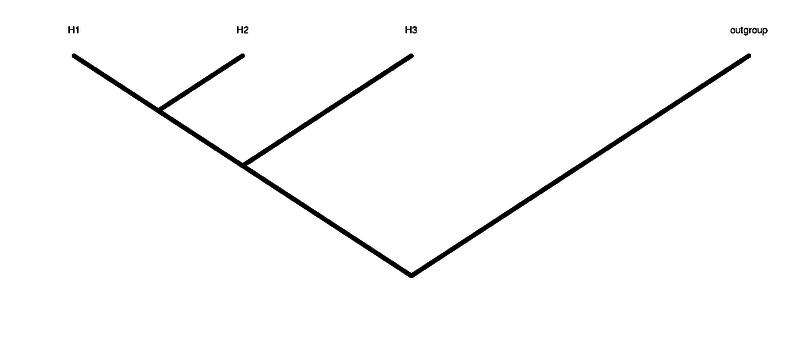
The ABBABABA (multipop) compute the abbababa test (aka D-statistic), that means testing for an ancient admixture (or wrong tree topology). Differently from ABBABABA (D_stat) multiple individuals for each one of the groups are allowed. As all methods in ANGSD we require that the header of the BAM files are the same.
This method has a publication.
- some of the options only works for the developmental version availeble from github
- if you use -rf to specify regions. These MUST appear in the same ordering as your fai file.
<classdiagram type="dir:LR">
[*.bam and/or *.cram| NGS genome datasets{bg:orange}]->[Sequence data|All bases or Random bases]
[Sequence data]->[Elaborate multiple genomes per population] [Elaborate multiple genomes per population]->[*.abbababa2|weighted ABBA and BABA counts file {bg:blue}] </classdiagram>
<classdiagram type="dir:LR"> [*.abbababa2|weighted ABBA and BABA counts file {bg:blue}]->estAvgError.R[D stat and Z scores{bg:blue}] </classdiagram>
Method
Brief Overview
> ./angsd -doAbbababa2
--------------
abcDstat2.cpp:
-doAbbababa2 0 run the abbababa analysis
-rmTrans 0 remove transitions
-blockSize 5000000 size of each block in bases
-anc (null) fasta file with outgroup
-sample 0 sample a single base in each individual
-maxDepth 100 max depth of each site allowed
-sizeFile (null) file with sizes of the populations
-enhance 0 only analyze sites where outgroup H4 is non poly
-Aanc 0 set H4 outgroup allele as A in each site
-useLast 0 1=use the last group of bam files as outgroup
This function will counts the number of ABBA and BABA sites of the 4-population trees that can be built from the data, where the outgroup is fixed.
Output
- 1)*.abbbababa2 (used for the 4-population test)
Each line represents a block with a chromsome name (Column 1), a start position (Column 2), an end postion (Column 3). Columns 4 and 5 are the numerator and denominator of the D-statistic for their specific block. Column 6 is the number of sites containing data in that block. The other 256 columns are the normalized counts of the 256 allele patterns between the 4 populations, starting from X0000=AAAA,X0001=AAAC,....,X3333=TTTT, with the correspondence 0=A,1=C,2=G,3=T. Every block is repeated a number of times corresponding to the trees that are built. This file is used as input for the R script estAvgError.R.
Options
- -doAbbababa2 1
take all bases at each position.
- -rmTrans [int]
0; use all reads (default), 1 Remove ancient transitions (important for ancient DNA)
- -blockSize [int]
Size of each block. Choose a number that is higher than the LD in the populations. For human 5Mb (5000000) is usually used.
- -anc [fileName.fa]
Include an outgroup in fasta format.
- -useLast [int]
1: use the last group of bam files as outgroup for the D-stat analysys. Default: 0 (use the fasta file as outgroup)
- -doCounts 1
use -doCounts 1 in order to count the bases at each sites after filters.
- -enhance [int]
1: use only sites where the reads for the outgroup has the same base for all reads.
- -sample [int]
1: sample only one base at each position for every individual 0: all bases at each position are used for the ABBABABA test
- -maxDepth [int]
allows for a maximum depth in each site to avoid overflow of the ABBA BABA counts. Default 100.
- -sizeFile [fileName]
file that specifies number of individuals in each population (more than 4 populations can be defined). If not provided, it is assumed that each population has only one individual.
- -Aanc [int]
1: H4 allele is A in each site.
In order to do fancy filtering of bases based on quality scores see the Allele counts options.
Tutorial for the ABBABABA (Multipop) test
This tutorial require having Samtools previously installed, and the library 'pracma' previously installed in R.
Prepare BAM and FASTA files
Download the latest version of angsd in your working folder from the github repository
https://github.com/ANGSD/angsd.git
Create symbolic links to angsd and the necessary R script
ln -s ./angsd/angsd ANGSD ln -s ./angsd/R/estAvgError.R DSTAT
Get 10 example .bam datasets, position them in the folder ./bams/ and create a file bam.filelist listing the pathnames of those datasets
wget http://popgen.dk/software/download/angsd/bams.tar.gz tar xf bams.tar.gz for i in bams/*.bam;do samtools index $i;done #index bam files ls bams/*.bam > bam.filelist rm bams.tar.gz #remove zipped file
This is how the file bam.filelist looks like
cat bam.filelist
bams/smallNA06985.mapped.ILLUMINA.bwa.CEU.low_coverage.20111114.bam bams/smallNA06994.mapped.ILLUMINA.bwa.CEU.low_coverage.20111114.bam bams/smallNA07000.mapped.ILLUMINA.bwa.CEU.low_coverage.20111114.bam bams/smallNA07056.mapped.ILLUMINA.bwa.CEU.low_coverage.20111114.bam bams/smallNA07357.mapped.ILLUMINA.bwa.CEU.low_coverage.20111114.bam bams/smallNA11829.mapped.ILLUMINA.bwa.CEU.low_coverage.20111114.bam bams/smallNA11830.mapped.ILLUMINA.bwa.CEU.low_coverage.20111114.bam bams/smallNA11831.mapped.ILLUMINA.bwa.CEU.low_coverage.20111114.bam bams/smallNA11832.mapped.ILLUMINA.bwa.CEU.low_coverage.20111114.bam bams/smallNA11840.mapped.ILLUMINA.bwa.CEU.low_coverage.20111114.bam
Download a fasta file for the chimpanzee. This is going to be used as the outgroup for the 4-population test.
wget http://popgen.dk/software/download/angsd/hg19ancNoChr.fa.gz mv hg19ancNoChr.fa.gz chimpHg19.fa.gz gunzip chimpHg19.fa.gz samtools faidx chimpHg19.fa #indexing the fasta file
Now, generate a fasta file for one of our 10 bam file. We assume such a genome has very high quality and we can use it as a reference for estimating error rates in others of our datasets.
./ANGSD -i bams/smallNA11840.mapped.ILLUMINA.bwa.CEU.low_coverage.20111114.bam -doFasta 1 -doCounts 1 -out perfectSampleCEU gunzip perfectSampleCEU.fa.gz samtools faidx perfectSampleCEU.fa
Generate files for the error correction
We will apply error correction to the group with 3 individuals, using "perfectSampleCEU" as high-quality reference genome. The population containing 3 individuals affected by transition error goes from line 6 to line 8 in the file bam.filelist. We select those individuals and write them in another file.
sed -n 6,8p bam.filelist > bamWithErrors.filelist
and then we use "doAncError" to generate the intermediate files that we will use later as input for the R script that calculates the D-statistic. "doAncError" applies the so called "perfect individual assumption", based on which error rates are estimate using a high quality genome (option -ref) and an outgroup (option -anc), both in fasta format. We have already prepared the two fasta files in our preparation phase.
./ANGSD -doAncError 1 -anc chimpHg19.fa -ref perfectSampleCEU.fa -out errorFile -bam bamWithErrors.filelist
4-population test
In this tutorial we perform the ABBABABA test on all the combinations of 4 populations amongst 6 populations of size 1,2,2,3,2,1 individuals, where the last population is fixed as outgroup (so that there are 30 possible combinations). The last population is represented by the fasta file defined with the option -anc, of which we enable the use as outgroup by the option -useLast 0. One can use the last population of .bam files as outgroup with the option -useLast 1. Create a file named sizeFile.size and write the size of each population (IT IS NECESSARY to define the size of the -anc outgroup population, that is always 1):
1 2 2 3 2 1
We decide to target three chromosomes, one of the three with loci between position 10Mb and 15Mb. Thus create a file called regions.txt in which is written
1: 5: 16:10000000-15000000
The output of ANGSD will show no data about chromosome 1. This happens when all blocks within that chromosome contained no data and therefore where not printed.
After running ANGSD to count ABBA and BABA combinations, we will call the R script who applies error correction to the ABBA and BABA allele combinations and produces the final output files.
./ANGSD -doAbbababa2 1 -bam bam.filelist -sizeFile sizeFile.size -doCounts 1 -out bam.Angsd -anc chimpHg19.fa -rf regions.txt -useLast 0 -minQ 20 -minMapQ 30 -p 1
The output file is bam.Angsd.abbbababa2 (used for the 4-population test) Each line represents a block with a chromsome name (Column 1) for one of the possible 30 trees (so each block is written on 30 lines), a start position (Column 2), an end postion (Column 3). Columns 4,5 and 6 are the numerator, denominator and number of sites analyzed in the block. The next 256 columns are the counted patterns of alleles in the tree, starting from X0000=AAAA,X0001=AAAC,....,X3333=TTTT, with the correspondence 0=A,1=C,2=G,3=T. This file is used as input for the R script estAvgError.R.
We run the R script specifying the error files for the population with 3 individuals. This is done defining the error files in each populations inside a text file (including a line for the outgroup population). If a population has no error file, it is sufficient to write NA. Create a file called errorList.error with written
NA NA NA ./errorFile.ancError NA NA
Create a file popNames.name with written
Population1 Population2 Population3 PopWithError Population4 Chimpanzee
Run the Rscript with the command
Rscript DSTAT angsdFile="bam.Angsd" out="result" sizeFile=sizeFile.size errFile=errorList.error nameFile=popNames.name
The script will show the calculated D statistic along with Z-score, Pvalues, Standard deviation and other quantities for all 30 4-populations trees. Note: If error correction is not needed, it is sufficient to avoid specifying any error file. If no names need to be provided, the script will assign Population_* as standard name. If no size file is provided, the script assigns 1 to each population. At least one between the name file and the size file is needed. It is possible to recycle the size file used in ANGSD.
The D-statistics and other informations are contained in four distinct files depending on the application of error correction and ancient transition removal. The files are named as follow:
D-statistic calculated WITHOUT Error Correction and WITHOUT Ancient Transition removal
D-statistic calculated WITH Error Correction and WITHOUT Ancient Transition removal
D-statistic calculated WITH Error Correction and WITH Ancient Transition removal
D-statistic calculated WITHOUT Error Correction and WITH Ancient Transition removal
Specifically, the values contained in the four files are: mean(D)=average D-stat, JK-D=jackknife estimate of the D-stat, V(JK-D)=variance of the D-stat, Z=Z score, pvalue=pvalue from the Z score, nABBA=number of ABBA patterns observed, nBABA=number of BABA patterns observed, nBlocks=number of blocks with observed data, H*=the names of the four populations for the specific tree. Note that the number of patterns might not be integer because of how ANGSD treats multiple genomes per populations.
Cite the method
@article{Soraggi2018,
author = {Soraggi, S. and Wiuf, C. and Albrechtsen, A.},
doi = {10.1534/g3.117.300192},
issn = {21601836},
journal = {G3: Genes, Genomes, Genetics},
number = {2},
title = {{Powerful inference with the D-statistic on low-coverage whole-genome data}},
volume = {8},
year = {2018}
}